Welcome to the latest post in the Graphs for Data Science substack. In this post, we explore the structure of networks using k-core decomposition, a unique way to probe the connectivity patterns of complex networks.
Real-world networks are typically large and hard to visualize, leading to the development of sophisticated methods and algorithms, such as degree correlations, network structure, and modularity, to explore their structure systematically. In this post, we continue this exploration by learning about k-core decomposition.
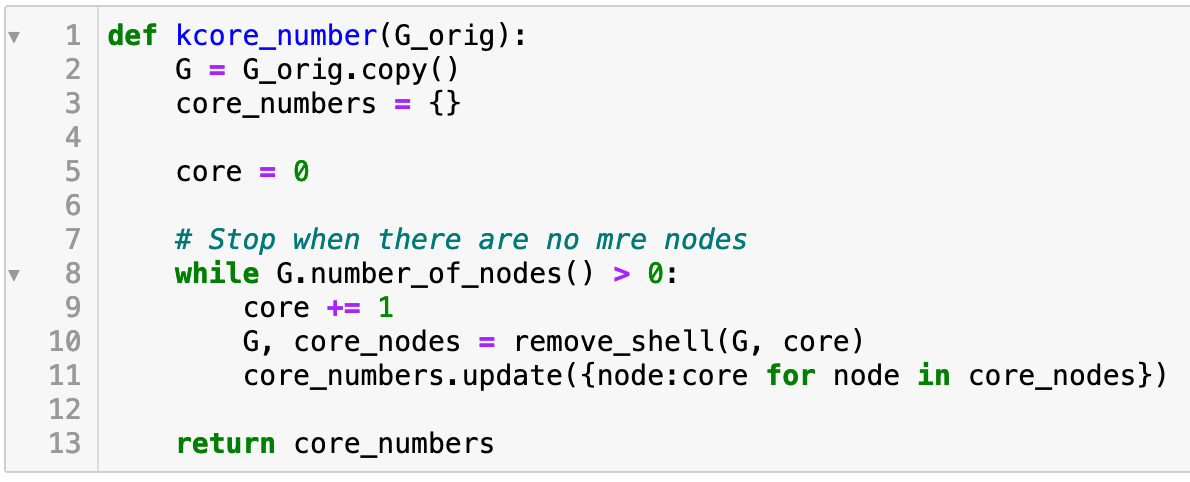
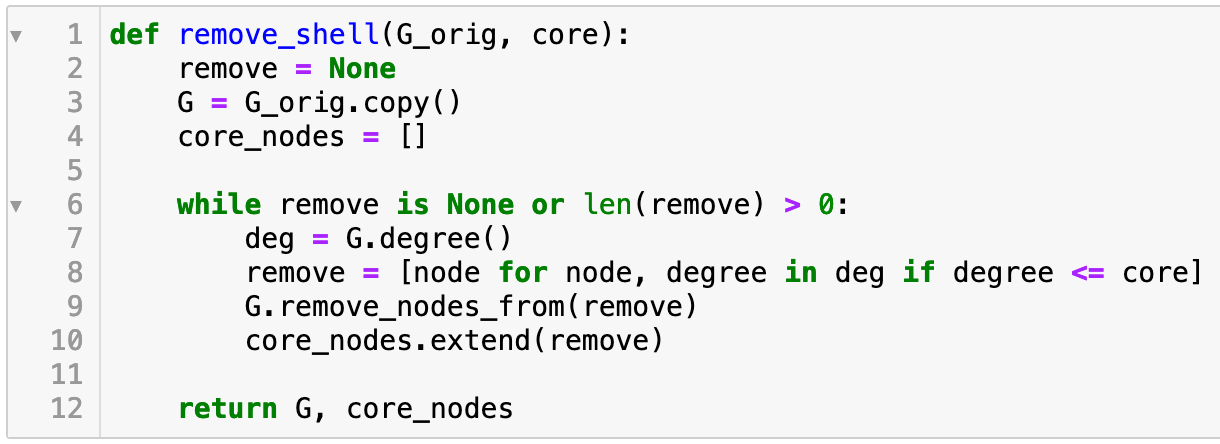
The k-core of a graph is the largest subgraph where every node has degree of at least k. The k-core can easily computed by recursively removing every node of degree less than k. In Python, this can done with:
As we move from one k-core to the next, the set of all removed nodes is known as the k-shell. If we apply this procedure to every value of the degree k and label each node by the k-shell to which it belongs, we obtain the k-core decomposition of the graph:
In the case of the Karate Club network, we obtain:
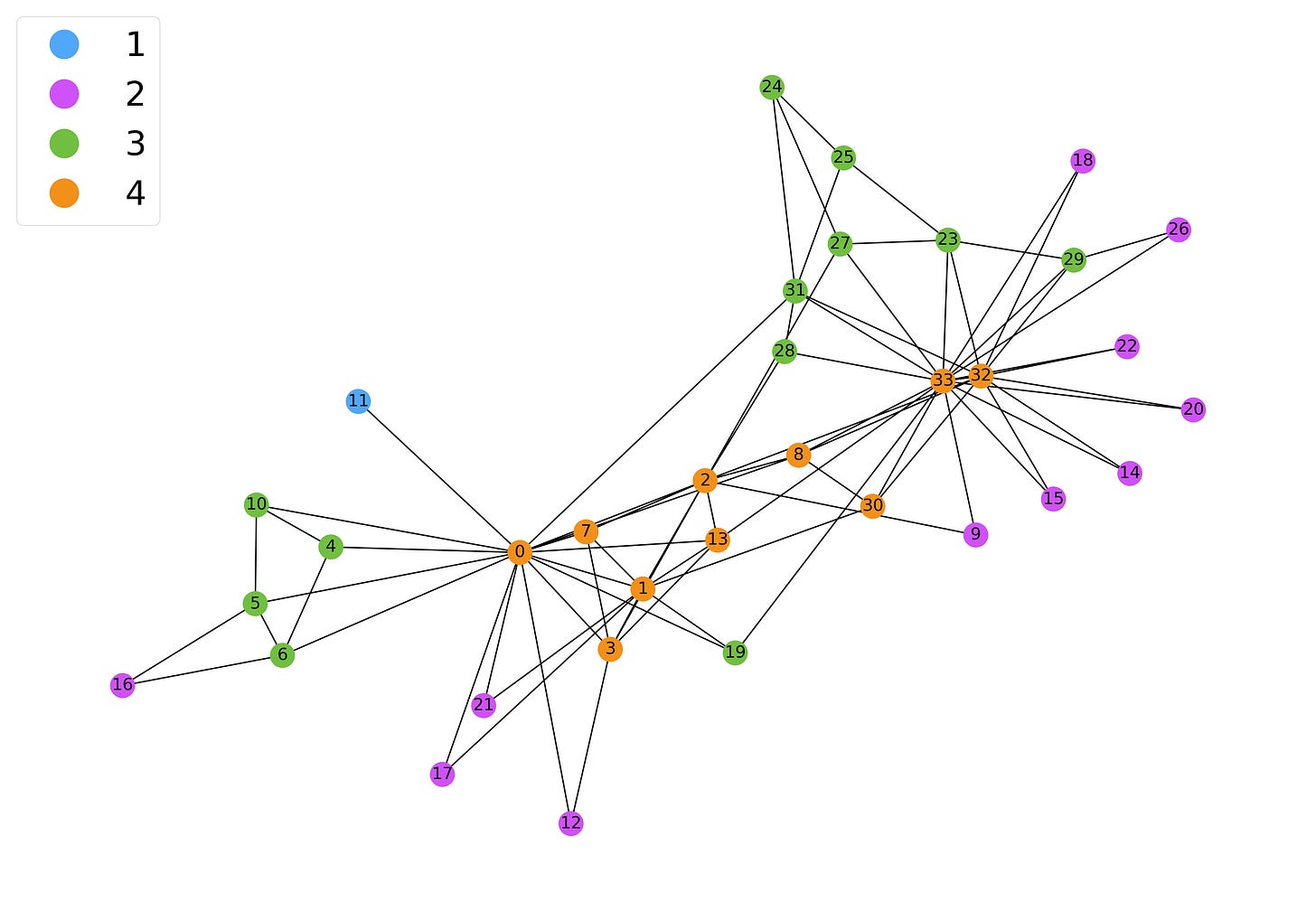
Naturally, the value of the k-core is higher the more central and well-connected a node is. There are two interesting edge cases we should consider.
The first case is that of a tree. Due to the recursive nature of the k-core decomposition algorithm, it is easy to see that every node in a tree has only a k-core of 1. The reason why is easy to understand: leaves always have degree one, so they get peeled off in the first pass. This, in turn, makes every node in the first level away from the leaves become leaves, so they get pruned in the second pass, and so on, until all nodes are removed.
In the opposite extreme, we have the case of a complete graph of N nodes. In this case, every node is connected to N-1 other nodes so the recursive procedure will fail to make any progress until it reaches the N-1 core, at which point it will remove all the nodes in the graph. In other words, a complete graph of size N only has nodes in the N-1 shell.
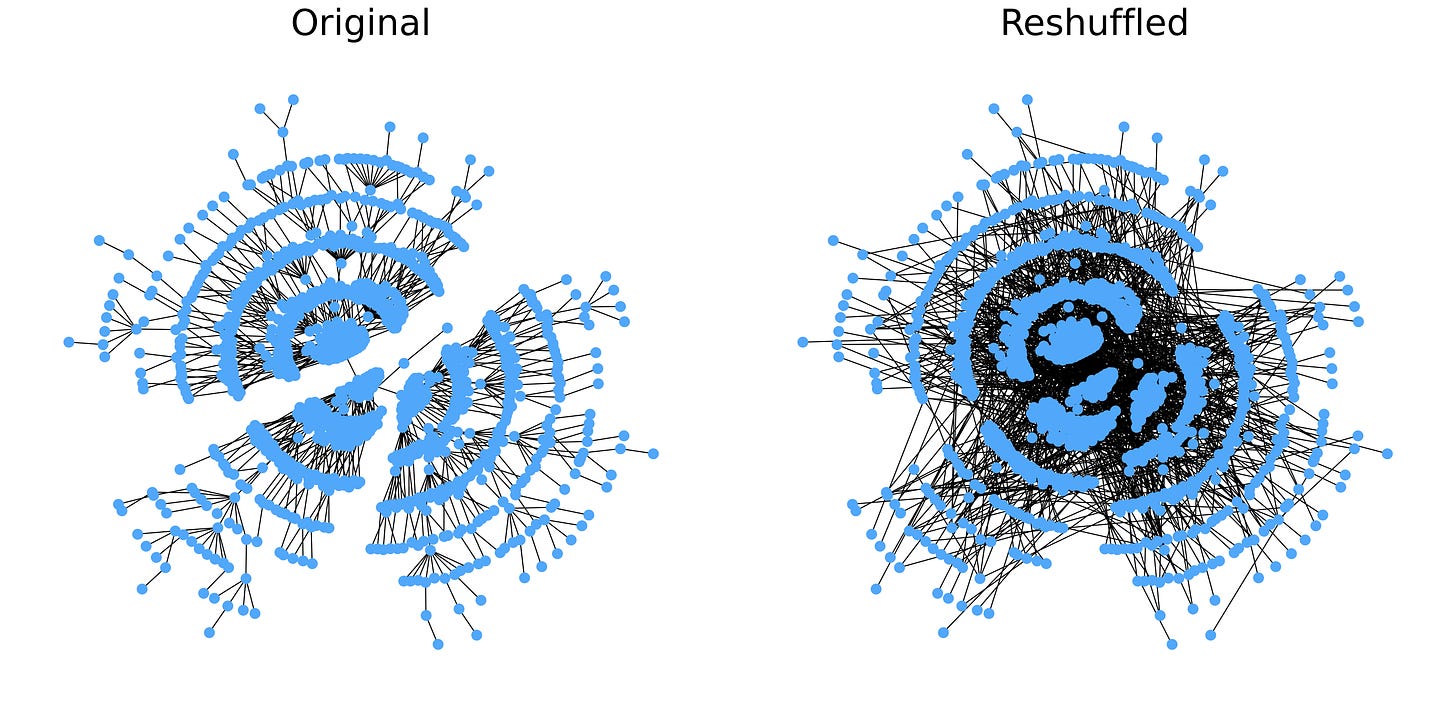
We can further refine our understanding of the k-core decomposition by looking at a simple Barabási-Albert Graph. If each node starts with degree one, the resulting graph is a tree with a broad-tailed degree distribution. We can break the tree structure of the graph by reshuffling all the edges while maintaining the degree of each node.
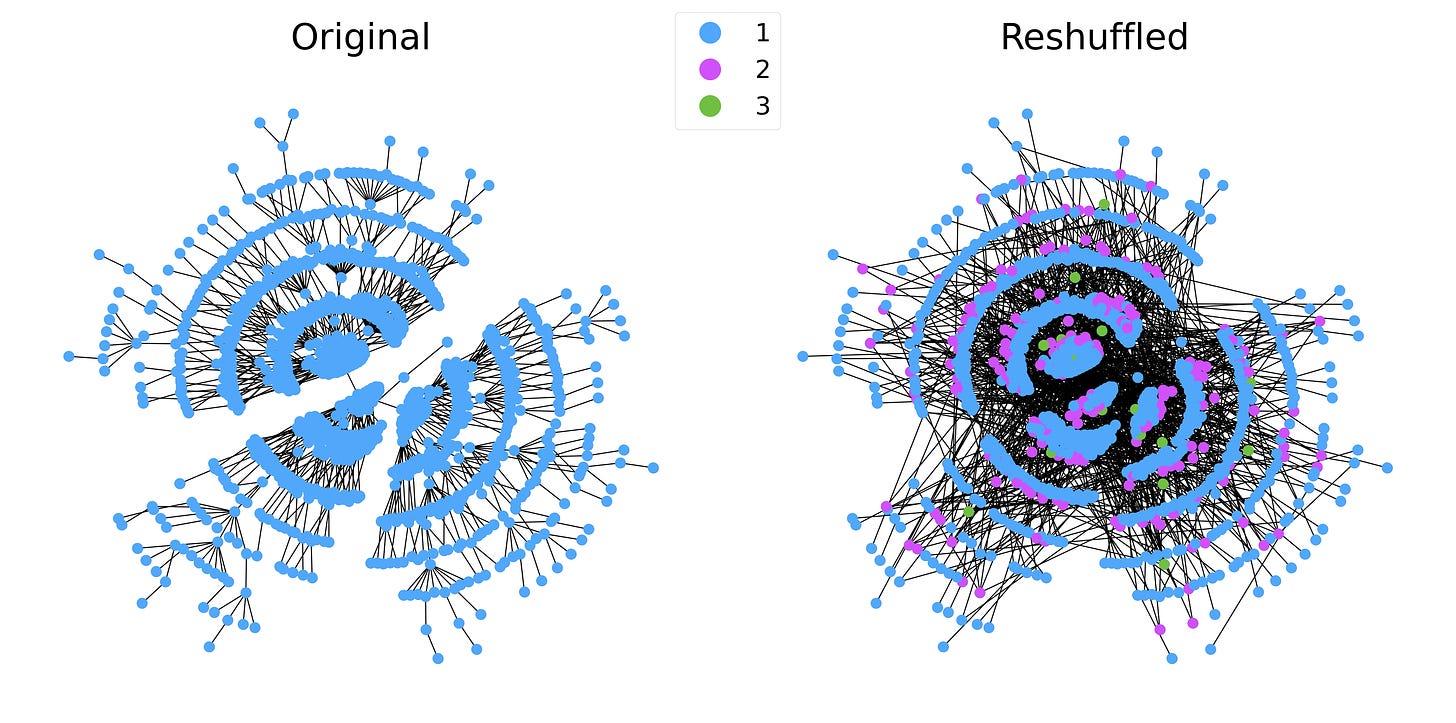
The reshuffled graph has a general structure that differs significantly from a tree. The k-core structure is also very different: the tree has every node in the 1-core, while the reshuffled graph has 3 k-cores with nodes in each core scattered across the entire graph.
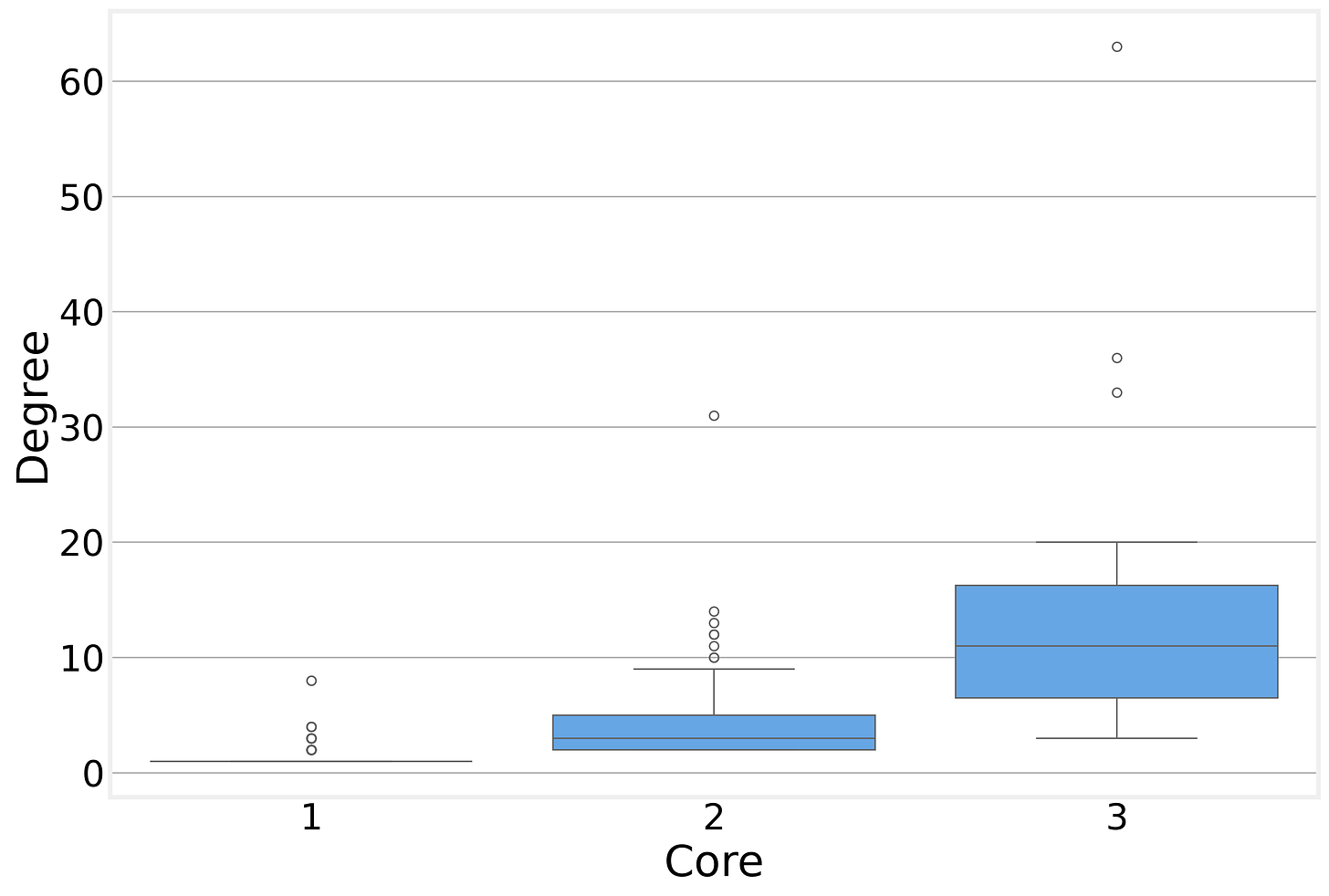
This also makes it clear that the degree of the node doesn’t need to match the core it belongs to. While the degree of each node belonging to a given k-shell is at least k, there is no upper limit. In our BA example, the maximum degree is over 60, while the network has a maximum k-core of just 3.
The k-core decomposition provides us with unique information about the way a network is connected and is one more tool we can use to explore the structure of networks.
We hope you enjoyed this post on the Graphs for Science Substack and look forward to hearing your thoughts. We hope you
You can find all the code for the analysis in this post in our companion GitHub Repository https://github.com/DataForScience/Graphs4Sci
And, of course, don’t forget to
this post with others who might be interested and encourage them to
so that they have access to the entire backlog of posts and be the first to know when a new a new article is posted.